Publications
2025
-
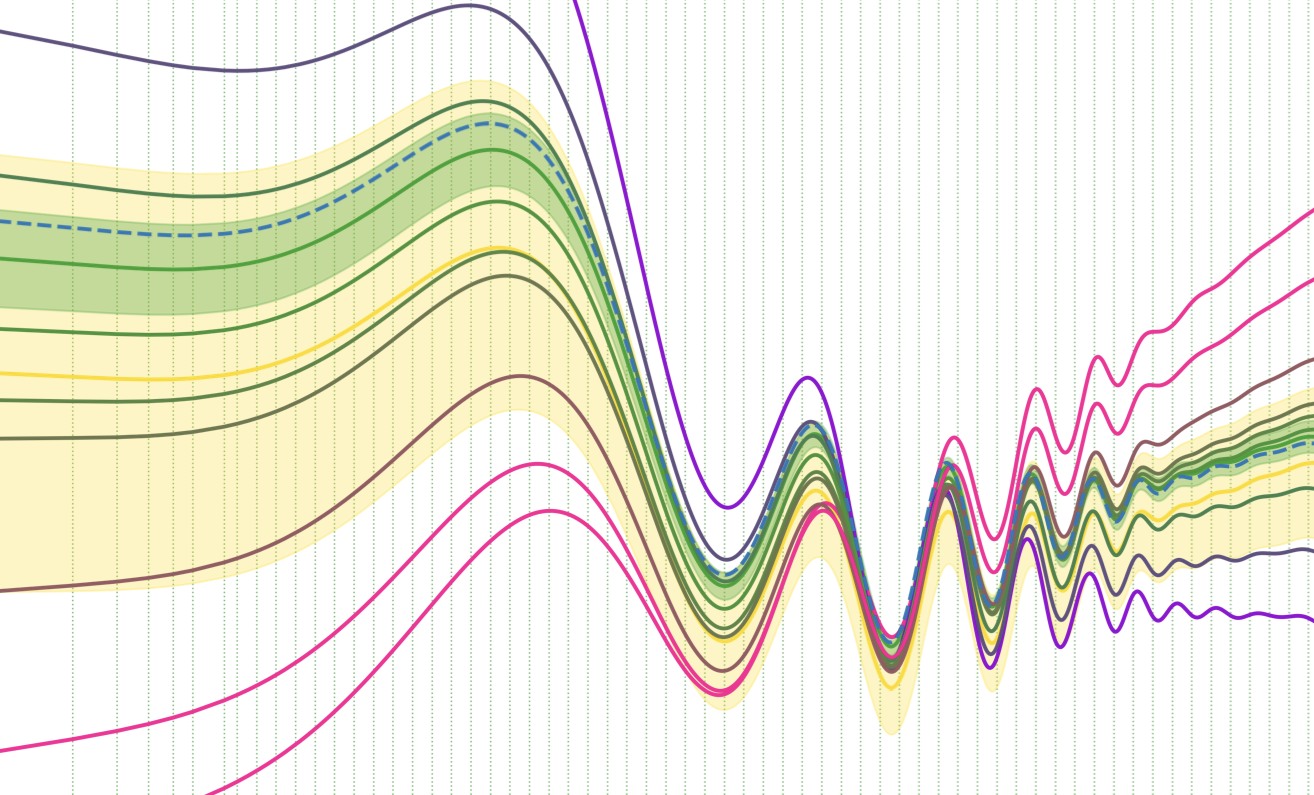 Diagnosing systematic effects using the inferred initial power spectrumTristan Hoellinger, and Florent LeclercqAstronomy & Astrophysics Jul 2025
Diagnosing systematic effects using the inferred initial power spectrumTristan Hoellinger, and Florent LeclercqAstronomy & Astrophysics Jul 2025Context. The next generation of galaxy surveys has the potential to substantially deepen our understanding of the Universe. This potential hinges on our ability to rigorously address systematic uncertainties. Until now, diagnosing systematic effects prior to inferring cosmological parameters has been out of reach in field-based implicit likelihood cosmological inference frameworks. Aims. As a solution, we aim to diagnose a variety of systematic effects in galaxy surveys prior to inferring cosmological parameters, using the inferred initial matter power spectrum. Methods. Our approach is built upon a two-step framework. First, we employed the simulator expansion for likelihood-free inference (SELFI) algorithm to infer the initial matter power spectrum, which we utilised to thoroughly investigate the impact of systematic effects. This investigation relies on a single set of N-body simulations. Second, we obtained a posterior on cosmological parameters via implicit likelihood inference, recycling the simulations from the first step for data compression. As a demonstration, we relied on a model of large-scale spectroscopic galaxy surveys that incorporates fully non-linear gravitational evolution with COmoving Lagrangian Acceleration (COLA) and simulates multiple systematic effects encountered in real surveys. Results. We provide a practical guide on how the SELFI posterior can be used to assess the impact of misspecified galaxy bias parameters, selection functions, survey masks, inaccurate redshifts, and approximate gravity models on the inferred initial matter power spectrum. We show that a subtly misspecified model can lead to a bias exceeding 2\ensuremathσ in the (\ensuremathΩ_m, \ensuremathσ_8) plane, which we are able to detect and avoid prior to inferring cosmological parameters. Conclusions. This framework has the potential to significantly enhance the robustness of physical information extraction from full forward models of large-scale galaxy surveys such as DESI, Euclid, and LSST.
@article{Hoellinger2025diagnosing, author = {{Hoellinger}, Tristan and {Leclercq}, Florent}, title = {{Diagnosing systematic effects using the inferred initial power spectrum}}, journal = {Astronomy {\&} Astrophysics}, keywords = {methods: statistical, cosmological parameters, large-scale structure of Universe, Cosmology and Nongalactic Astrophysics, Instrumentation and Methods for Astrophysics}, year = {2025}, month = jul, volume = {699}, eid = {A224}, pages = {A224}, doi = {10.1051/0004-6361/202453416}, archiveprefix = {arXiv}, eprint = {2412.04443}, primaryclass = {astro-ph.CO}, adsurl = {https://ui.adsabs.harvard.edu/abs/2025A&A...699A.224H}, adsnote = {Provided by the SAO/NASA Astrophysics Data System}, } -
 A vorticity confinement correction for discontinuous Galerkin schemes applied to fluid flow problemsTristan Hoellinger, Jean-Baptiste Chapelier, and Lucas ManuecoInt. Journal of Numerical Methods for Heat & Fluid Flow Jan 2025
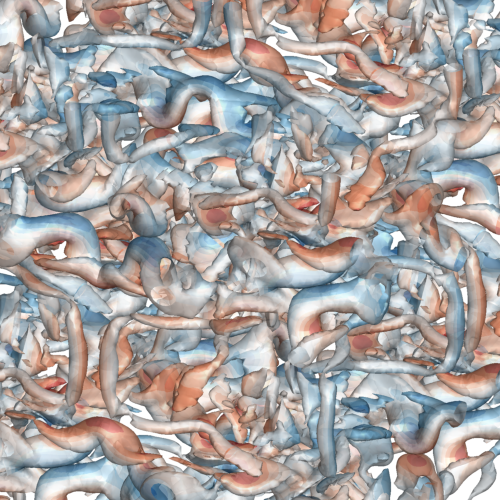
A vorticity confinement correction for discontinuous Galerkin schemes applied to fluid flow problemsTristan Hoellinger, Jean-Baptiste Chapelier, and Lucas ManuecoInt. Journal of Numerical Methods for Heat & Fluid Flow Jan 2025Purpose Vortices are ubiquitous in fluid flows, yet accurately preserving their dynamics in numerical simulations over extended timescales and distances remains challenging. Discretisation schemes often introduce artificial dissipation exceeding physical viscosity, flattening vortical structures and disrupting their delicate dynamics. This paper aims to introduce a vorticity confinement correction for discontinuous Galerkin discretisation methods, designed to enhance the fidelity of direct numerical simulations of unsteady, vortex-dominated flows at arbitrarily high orders of accuracy. Design/methodology/approach This approach builds upon the classical VC2 correction for finite volume schemes, incorporating an additional volumetric correction specific to the discontinuous Galerkin discretisation. Findings This paper shows through an isentropic vortex transport problem that the vorticity confinement correction substantially improves vorticity conservation, evaluated here at second- and third-order spatial accuracies. This paper further demonstrates a significant gain in preserving both large and small vortices in three-dimensional transitional turbulence at second-order spatial accuracy. Originality/value To the best of the authors’ knowledge, this is the first time a Vorticity Confinement method is developed and evaluated in the context of discontinuous Galerkin schemes.
@article{Hoellinger2025, author = {Hoellinger, Tristan and Chapelier, Jean-Baptiste and Manueco, Lucas}, title = {A vorticity confinement correction for discontinuous Galerkin schemes applied to fluid flow problems}, journal = {Int. Journal of Numerical Methods for Heat {\&} Fluid Flow}, year = {2025}, month = jan, day = {01}, publisher = {Emerald Publishing Limited}, volume = {ahead-of-print}, number = {ahead-of-print}, doi = {10.1108/HFF-11-2024-0854}, url = {https://doi.org/10.1108/HFF-11-2024-0854}, }
2023
-
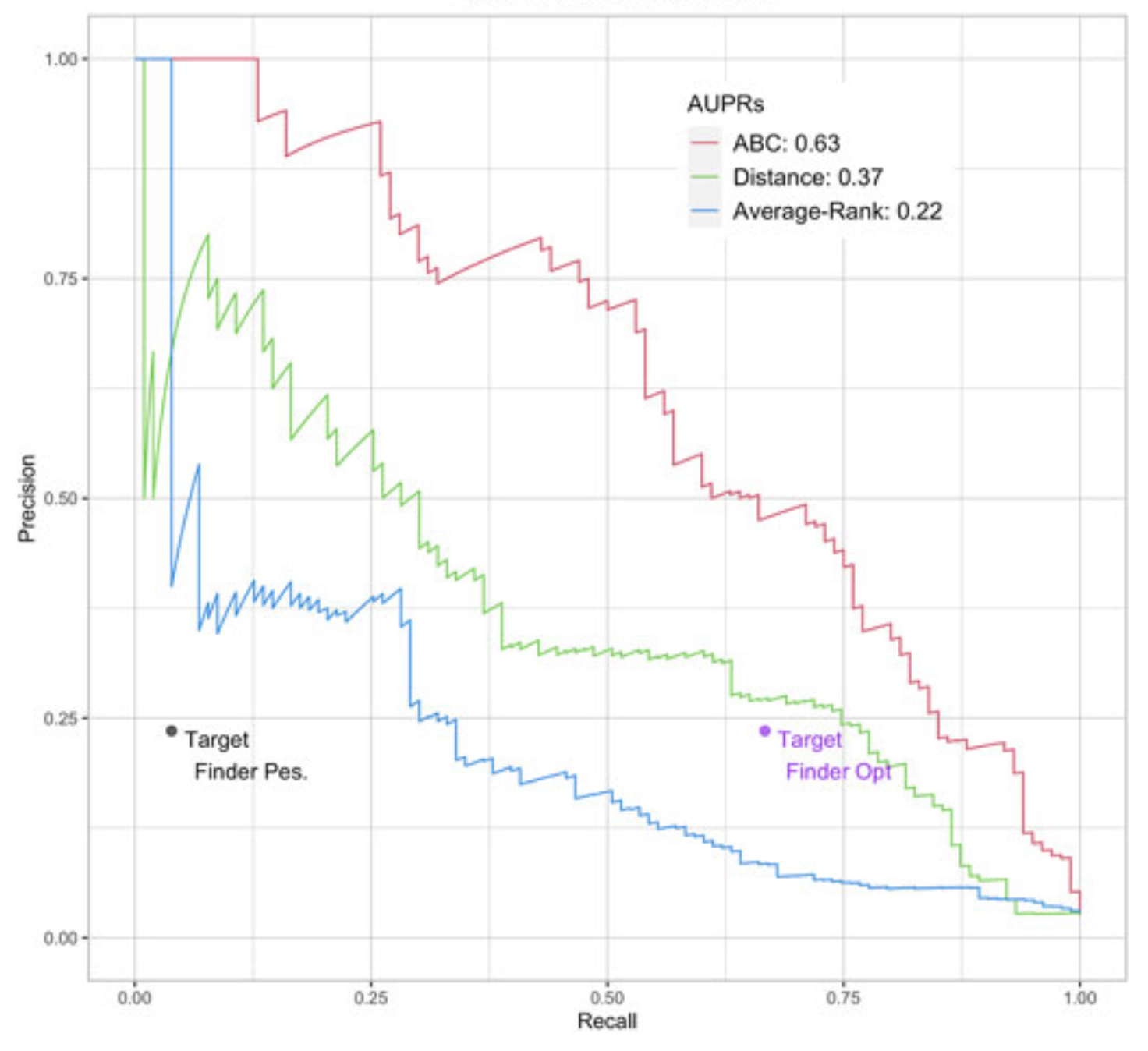 Enhancer/gene relationships: need for more reliable genome-wide reference setsTristan Hoellinger, Camille Mestre, Hugues Aschard, and 4 more authorsFrontiers in Bioinformatics Jan 2023
Enhancer/gene relationships: need for more reliable genome-wide reference setsTristan Hoellinger, Camille Mestre, Hugues Aschard, and 4 more authorsFrontiers in Bioinformatics Jan 2023Differences in cells’ functions arise from differential activity of regulatory elements, including enhancers. Enhancers are cis-regulatory elements that cooperate with promoters through transcription factors (TF) to activate the expression of one or several genes by getting physically close to them in the 3D space of the nucleus. There is increasing evidence that genetic variants associated with common diseases are enriched in enhancers active in cell types relevant to these diseases. Identifying the enhancers associated with genes and conversely, the sets of genes activated by each enhancer (the so-called enhancer/gene or E/G relationships) across cell types, can help understanding the genetic mechanisms underlying human diseases. There are three broad approaches for the genome-wide identification of E/G relationships in a cell type: (1) genetic link methods or eQTL, (2) functional link methods based on 1D functional data such as open chromatin, histone mark or gene expression and (3) spatial link methods based on 3D data such as HiC. Since (1) and (3) are costly, the current strategy is to develop functional link methods and to use data from (1) and (3) as reference to evaluate them. However, there is still no consensus on the best functional link method to date, and method comparison remain seldom. Here, we compared the relative performances of three recent methods for the identification of18 enhancer-gene links, TargetFinder, Average-Rank, and the ABC model, using the three19 latest benchmarks from the field: a reference that combines 3D and eQTL data, called BENGI,20 and two genetic screening references, called CRiFF and CRiSPRi. Overall, none of the three21 methods performed best on the three references. CRiFF and CRISPRi reference sets are likely22 more reliable, but CRiFF is not genome-wide and CRiFF and CRISPRi are mostly available on23 the K562 cancer cell line. The BENGI reference set is genome-wide but likely contains many false24 positives. This study therefore calls for new reliable and genome-wide E/G reference data rather25 than new functional link E/G identification methods.
@article{hoellinger2023, title = {Enhancer/gene relationships: need for more reliable genome-wide reference sets}, author = {Hoellinger, Tristan and Mestre, Camille and Aschard, Hugues and Le Goff, Wilfried and Foissac, Sylvain and Faraut, Thomas and Djebali, Sarah}, journal = {Frontiers in Bioinformatics}, year = {2023}, volume = {3}, section = {Insights in Bioinformatics}, doi = {10.3389/fbinf.2023.1092853}, }
2020
-
 Data-Driven Simulation for Augmented SurgeryAndrea Mendizabal, Eleonora Tagliabue, Tristan Hoellinger, and 3 more authorsAdvanced Structured Materials, Springer Jan 2020
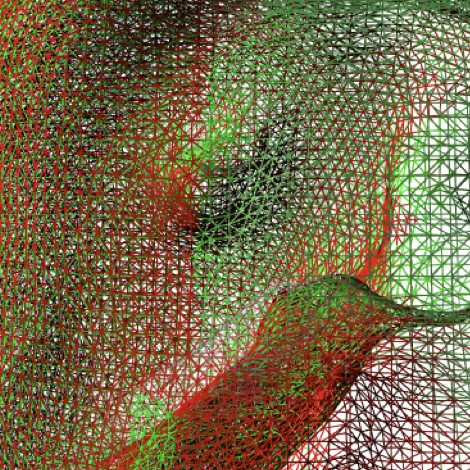
Data-Driven Simulation for Augmented SurgeryAndrea Mendizabal, Eleonora Tagliabue, Tristan Hoellinger, and 3 more authorsAdvanced Structured Materials, Springer Jan 2020To build an augmented view of an organ during surgery, it is essential to have a biomechanical model with appropriate material parameters and boundary conditions, able to match patient specific properties. Adaptation to the patient’s anatomy is obtained by exploiting the image-rich context specific to our application domain. While information about the organ shape, for instance, can be obtained preoperatively, other patient-specific parameters can only be determined intraoperatively. To this end, we are developing data-driven simulations, which exploit information extracted from a stream of medical images. Such simulations need to run in realtime. To this end we have developed dedicated numerical methods, which allow for real-time computation of finite element simulations.
@article{Mendizabal2020, title = {Data-Driven Simulation for Augmented Surgery}, author = {Mendizabal, Andrea and Tagliabue, Eleonora and Hoellinger, Tristan and Brunet, Jean-Nicolas and Nikolaev, Sergei and Cotin, St{\'e}phane}, journal = {Advanced Structured Materials, Springer}, editor = {Abali, Bilen Emek and Giorgio, Ivan}, booktitle = {Developments and Novel Approaches in Biomechanics and Metamaterials}, year = {2020}, publisher = {Springer International Publishing}, address = {Cham}, pages = {71--96}, isbn = {978-3-030-50464-9}, doi = {10.1007/978-3-030-50464-9_5}, url = {https://doi.org/10.1007/978-3-030-50464-9_5}, }